Overview
- Cayman has made a SARS-CoV-2 (human) neutralizing recombinant antibody which disrupts the S1 RBD-ACE2 interaction.
- Surface plasmon resonance (SPR) was used to evaluate the recombinant neutralizing antibody’s ability to block binding of SARS-CoV-2 S1 receptor binding domain (RBD) variants to human angiotensin-converting enzyme 2 (hACE2).
- Cryo-electron microscopy (cryo-EM) single-particle analysis was used to determine the structure of intrinsically difficult-to-crystallize, >0.5 MDa SARS-CoV-2 spike:Fab complex.
- This case study shows the structure and conformational analysis of the SARS-CoV-2 spike glycoprotein trimer in complex with a Fab derived from a recombinant SARS-CoV-2 neutralizing antibody. Cayman’s engineered recombinant antibody could effectively block the interaction of hACE2 with S1 RBD wild-type and Alpha variants better than that with the Beta variant.
- These structural results from cryo-EM and SPR studies could aid in the development of vaccines and therapeutics for the SARS-CoV-2 variants.
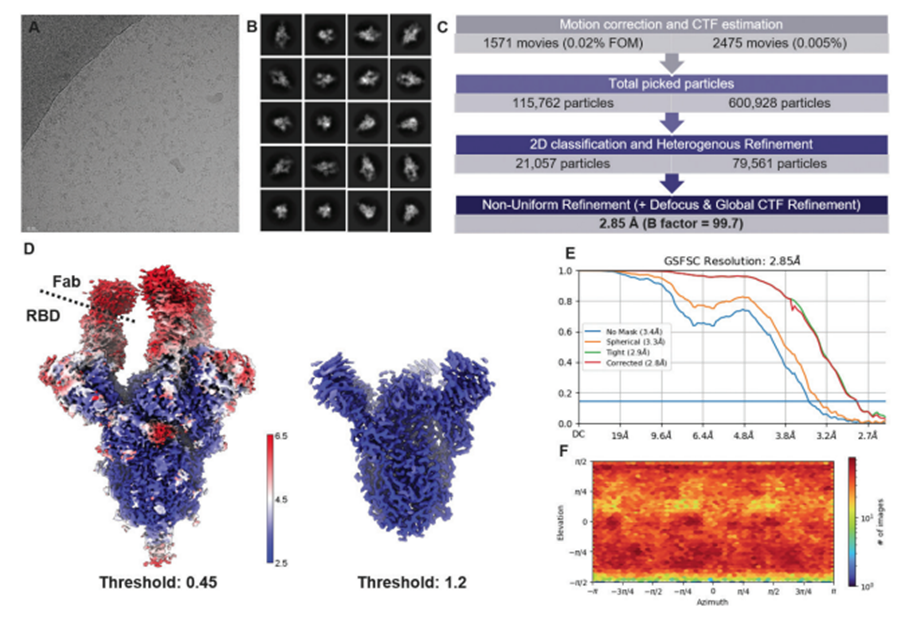
Figure 1. Cryo-EM single particle analysis of 6P-stabilized SARS-CoV-2 pre-fusion spike ectodomain in complex with a Fab.
Featured Products
SARS-CoV-2 (human) Neutralizing Recombinant Antibody
Disrupts the S1 RBD-ACE2 interaction and can be used for ELISA, immunofluorescence (IF), and Western blot (WB) applications. The antibody recognizes the SARS-CoV-2 spike glycoprotein at approximately 65 kDa.
SARS-CoV-2 Spike S1 RBD-ACE2 Binding Cellular Imaging Assay Kit
A sensitive and unique imaging platform for identifying novel modulators of SARS-CoV-2 spike S1 RBD and ACE2 binding. This assay uses Cayman’s proprietary reverse transfection platform to express human ACE2 on the surface of HEK293T/17 cells for binding by a recombinant rabbit Fc-tagged SARS-CoV-2 spike S1 RBD protein as a model of SARS-CoV-2 cellular infection.
